import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
def draw_box(ax, xy, w, h, text, color="#4A90D9", fontsize=9, text_color="white"):
rect = mpatches.FancyBboxPatch(xy, w, h, boxstyle="round,pad=0.06",
facecolor=color, edgecolor="black", linewidth=1.2)
ax.add_patch(rect)
ax.text(xy[0] + w/2, xy[1] + h/2, text, ha="center", va="center",
fontsize=fontsize, fontweight="bold", color=text_color)
def draw_arrow(ax, start, end, **kwargs):
ax.annotate("", xy=end, xytext=start,
arrowprops=dict(arrowstyle="-|>", color="black", lw=1.5, **kwargs))
fig, ax = plt.subplots(figsize=(12, 4))
ax.set_xlim(-0.5, 15)
ax.set_ylim(-1.5, 4.0)
ax.set_aspect("equal")
ax.axis("off")
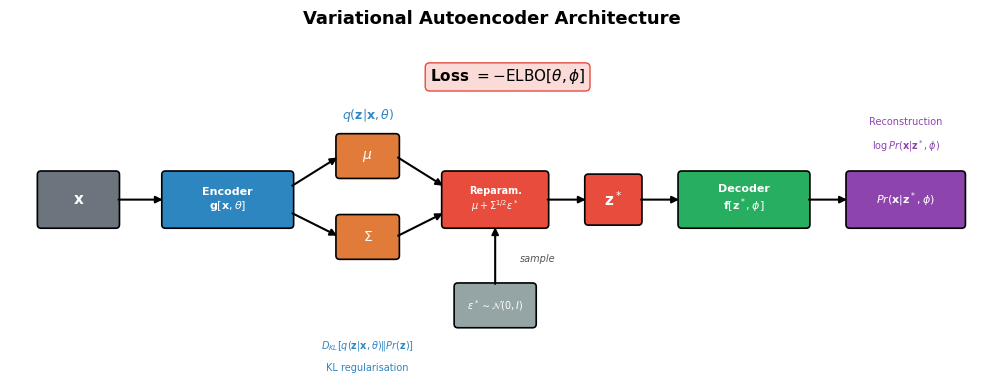
ax.set_title("Variational Autoencoder Architecture", fontsize=13, fontweight="bold", pad=10)
# Input x
draw_box(ax, (0, 1.0), 1.2, 0.8, r"$\mathbf{x}$", color="#6C757D", fontsize=11)
# Encoder
draw_box(ax, (2.0, 1.0), 2.0, 0.8, "Encoder\n$\\mathbf{g}[\\mathbf{x}, \\theta]$",
color="#2E86C1", fontsize=8)
draw_arrow(ax, (1.2, 1.4), (2.0, 1.4))
# mu and Sigma outputs
draw_box(ax, (4.8, 1.8), 0.9, 0.6, r"$\mu$", color="#E07B39", fontsize=10)
draw_box(ax, (4.8, 0.5), 0.9, 0.6, r"$\Sigma$", color="#E07B39", fontsize=10)
draw_arrow(ax, (4.0, 1.6), (4.8, 2.1))
draw_arrow(ax, (4.0, 1.2), (4.8, 0.8))
# q(z|x,theta) label
ax.text(5.25, 2.7, r"$q(\mathbf{z}|\mathbf{x}, \theta)$", ha="center", fontsize=9,
style="italic", color="#2E86C1")
# Reparameterization
draw_box(ax, (6.5, 1.0), 1.6, 0.8, "Reparam.\n$\\mu + \\Sigma^{1/2}\\epsilon^*$",
color="#E74C3C", fontsize=7)
draw_arrow(ax, (5.7, 2.1), (6.5, 1.6))
draw_arrow(ax, (5.7, 0.8), (6.5, 1.2))
# Epsilon noise
draw_box(ax, (6.7, -0.6), 1.2, 0.6, "$\\epsilon^* \\sim \\mathcal{N}(0,I)$",
color="#95A5A6", fontsize=7)
draw_arrow(ax, (7.3, 0.0), (7.3, 1.0))
ax.text(7.7, 0.4, "sample", fontsize=7, style="italic", color="#555")
# z*
draw_box(ax, (8.8, 1.05), 0.8, 0.7, r"$\mathbf{z}^*$", color="#E74C3C", fontsize=11)
draw_arrow(ax, (8.1, 1.4), (8.8, 1.4))
# Decoder
draw_box(ax, (10.3, 1.0), 2.0, 0.8, "Decoder\n$\\mathbf{f}[\\mathbf{z}^*, \\phi]$",
color="#27AE60", fontsize=8)
draw_arrow(ax, (9.6, 1.4), (10.3, 1.4))
# Reconstruction
draw_box(ax, (13.0, 1.0), 1.8, 0.8,
"$Pr(\\mathbf{x}|\\mathbf{z}^*, \\phi)$",
color="#8E44AD", fontsize=8)
draw_arrow(ax, (12.3, 1.4), (13.0, 1.4))
# Loss function label at top
ax.text(7.5, 3.3, r"Loss $= -\mathrm{ELBO}[\theta, \phi]$",
ha="center", fontsize=11, fontweight="bold",
bbox=dict(boxstyle="round,pad=0.3", facecolor="#FADBD8", edgecolor="#E74C3C"))
# Annotations for loss terms
ax.text(13.9, 2.2, r"$\log Pr(\mathbf{x}|\mathbf{z}^*, \phi)$",
ha="center", fontsize=7, color="#8E44AD", style="italic")
ax.text(13.9, 2.6, "Reconstruction", ha="center", fontsize=7, color="#8E44AD")
ax.text(5.25, -1.0, r"$D_{KL}[q(\mathbf{z}|\mathbf{x},\theta) \| Pr(\mathbf{z})]$",
ha="center", fontsize=7, color="#2E86C1", style="italic")
ax.text(5.25, -1.35, "KL regularisation", ha="center", fontsize=7, color="#2E86C1")
fig.tight_layout()
plt.show()